Reactivity and Specificity of RNase T1, RNase A, and RNase H toward Oligonucleotides of RNA Containing 8-Oxo-7,8-dihydroguanosine | Biochemistry
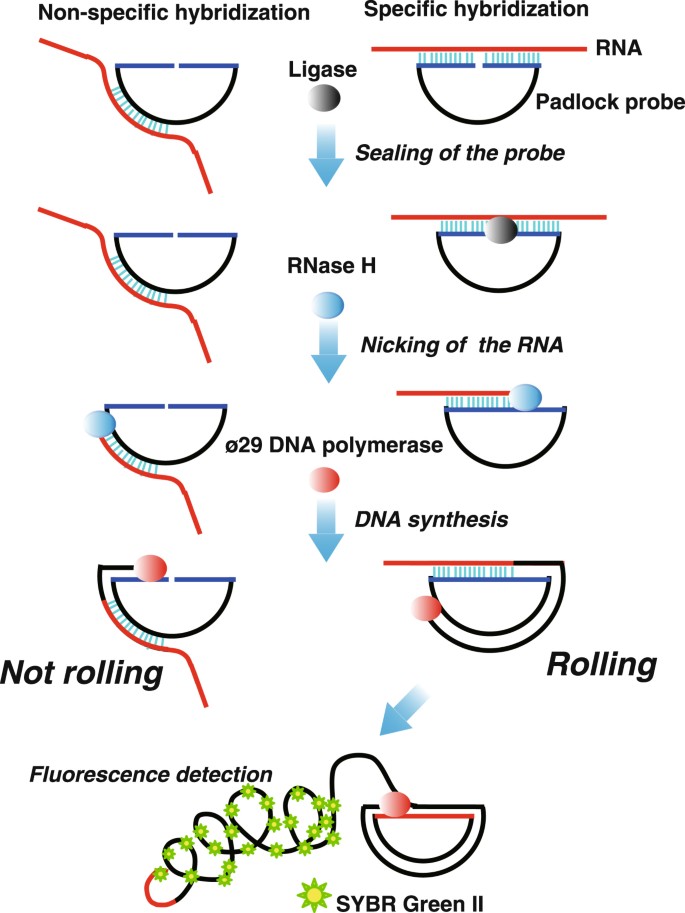
RNase H-assisted RNA-primed rolling circle amplification for targeted RNA sequence detection | Scientific Reports

Quantitative analysis of miRNAs using SplintR ligase-mediated ligation of complementary-pairing probes enhanced by RNase H (SPLICER)-qPCR: Molecular Therapy - Nucleic Acids

Characterization and Sequence Mapping of Large RNA and mRNA Therapeutics Using Mass Spectrometry | Analytical Chemistry
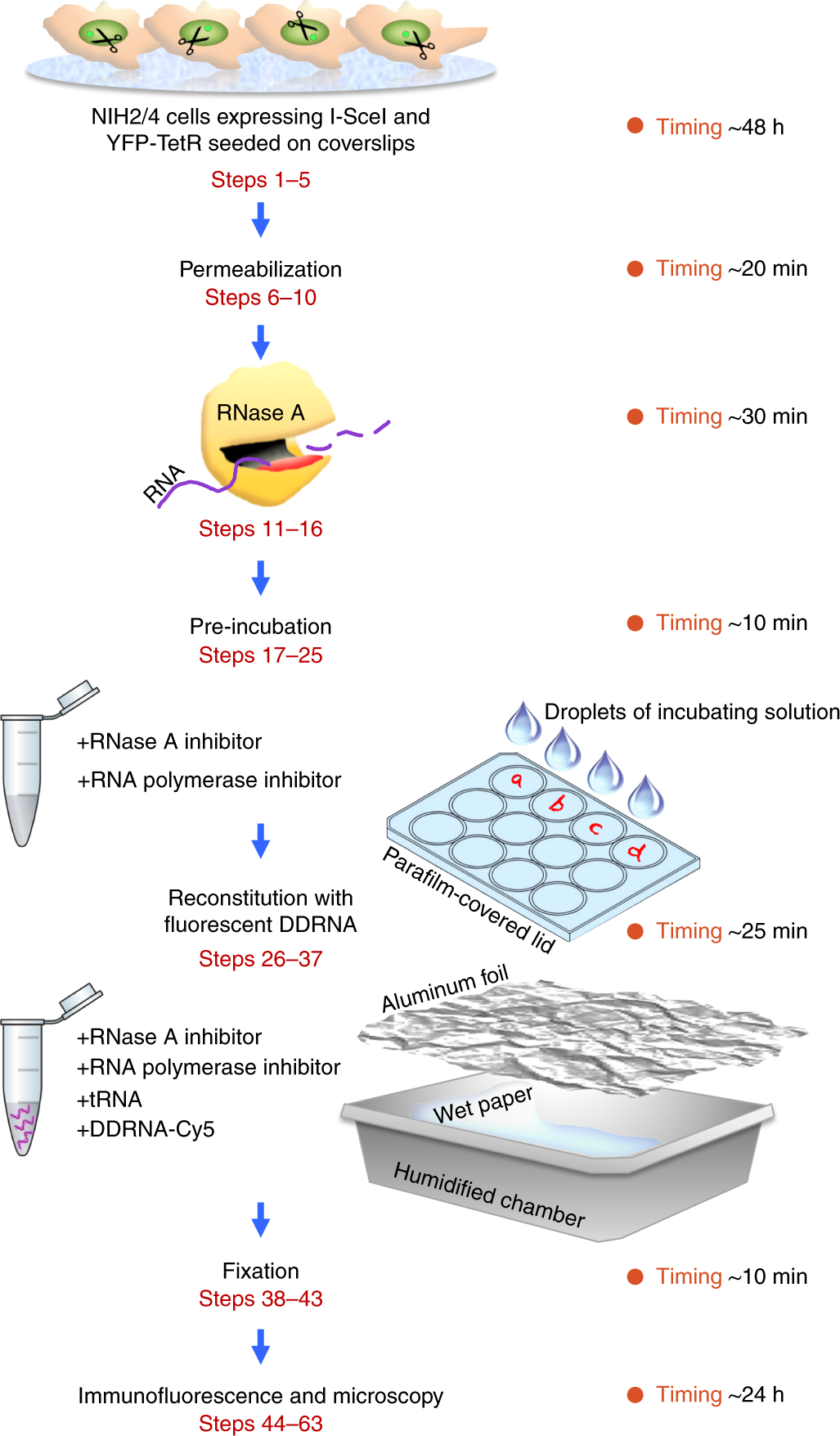
RNase A treatment and reconstitution with DNA damage response RNA in living cells as a tool to study the role of non-coding RNA in the formation of DNA damage response foci

MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein